Marine Laboratory
Island Evolution Lab
Island Evolution Lab
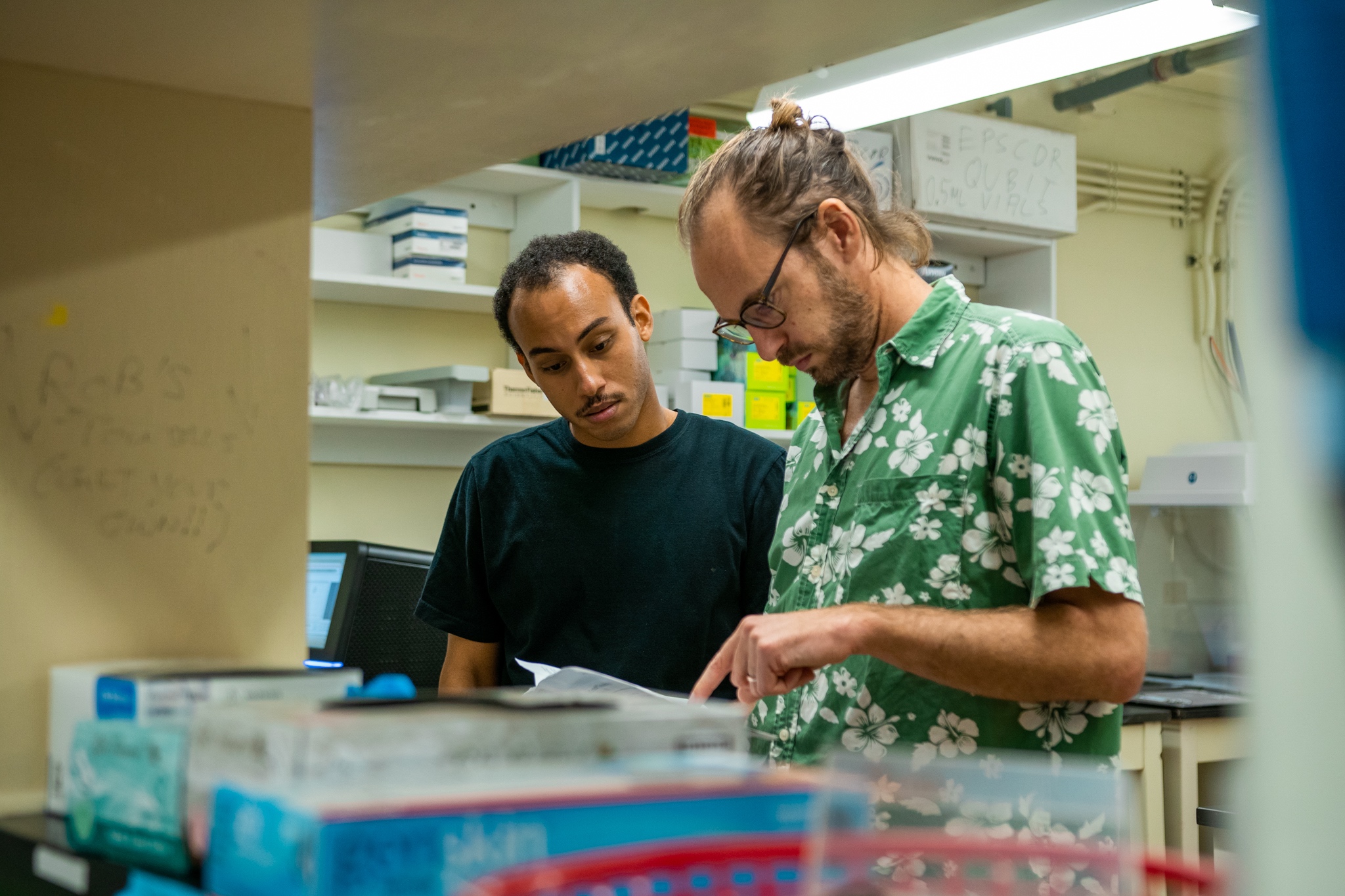
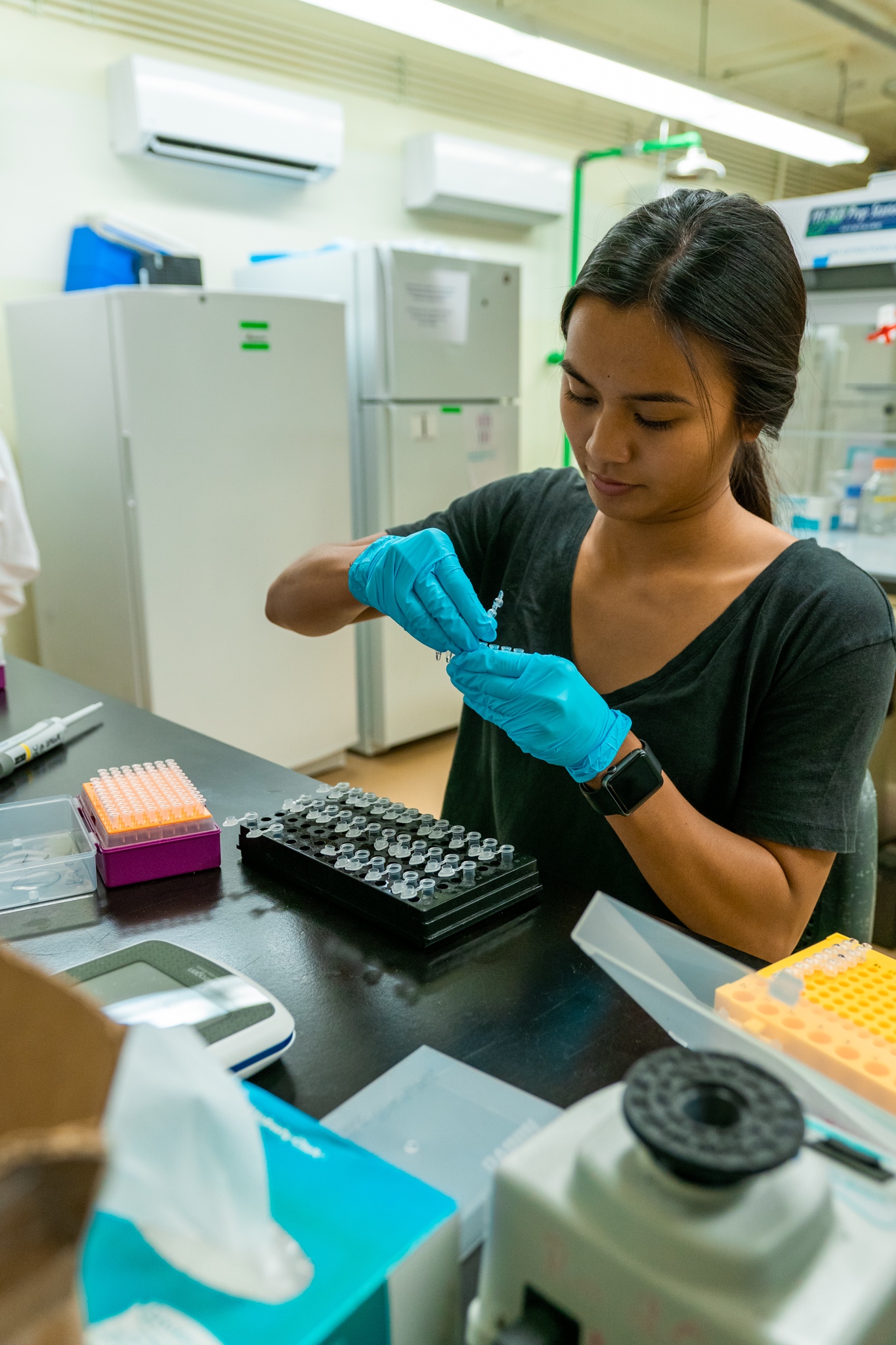
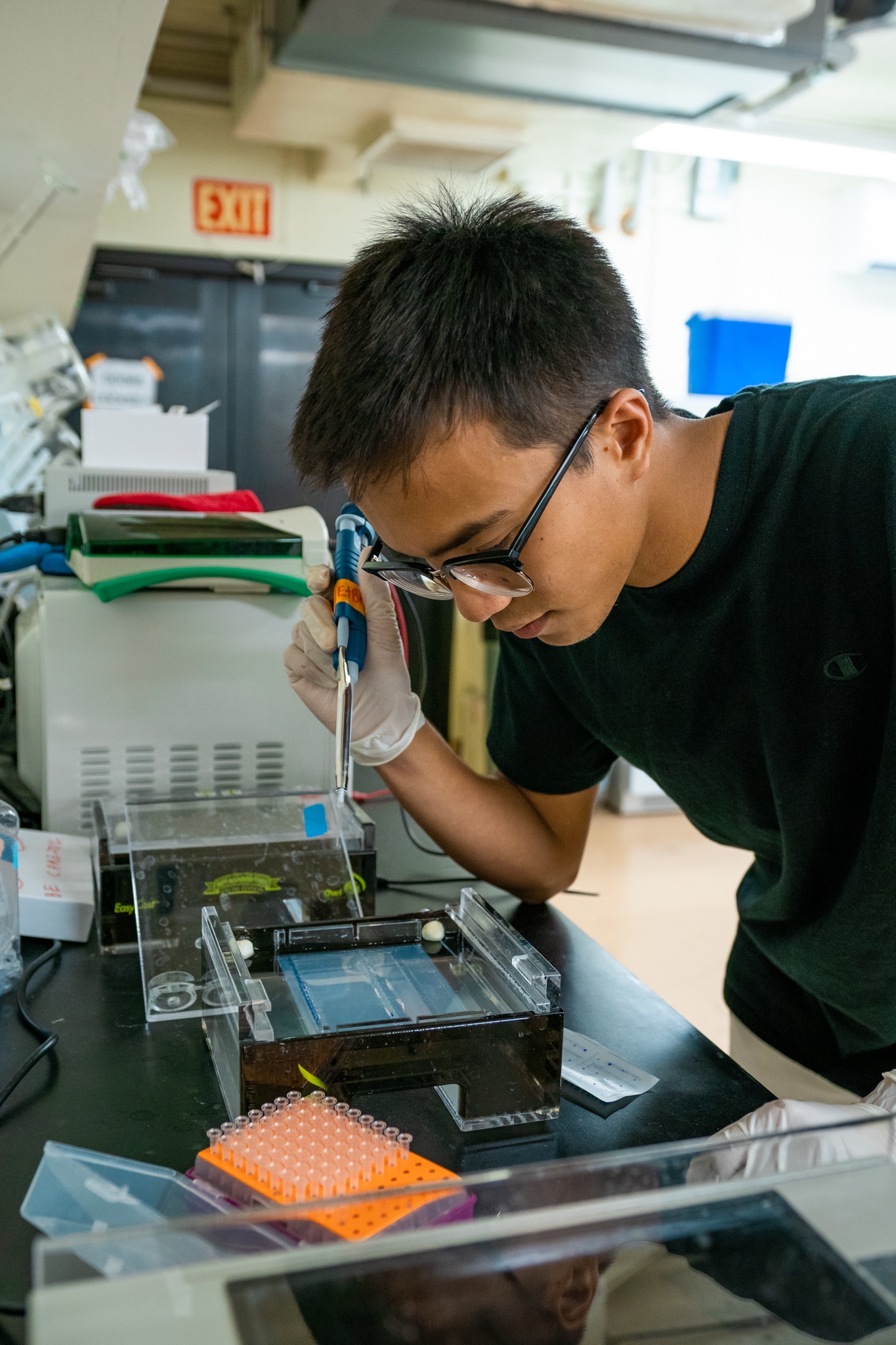
The Island Evolution Lab at the UOG Marine Lab is a group of researchers and students interested in basic and applied evolutionary question in island settings. We are using genetic and genomic approaches (RAD-Seq, RNA-Seq and Genome sequencing) in combination with field work, collections, museums, and experimental manipulations to address original questions in population genetics, phylogenetics, phylogeography, molecular ecology and genomics.
The Island Evolution Lab is partially funded by the National Science Foundation (NSF) EPSCoR Guam Ecosystems Collaboratorium program, NOAA's Coral Reef Conservation Program, the National Fish and Wildlife Foundation (NFWF) and the Government of Guam.
The Island Evolution Lab actively support and promotes diversity in science, research and education.

We are always looking for graduate and undergraduate students who are interested and passionate about evolution, genomics and population genetics!
Please contact us islandevolutionlab@gmail.com
Follow us on Twitter @Island_Evo_Lab (e.g. here in the column on the side -->
And on Instagram @islandevolutionlab (e.g. below)
Our instagram feed:
IEL Field work
A special feature about our lab on Guam's local TV station Kuam TV in April 2019 (also on youtube)
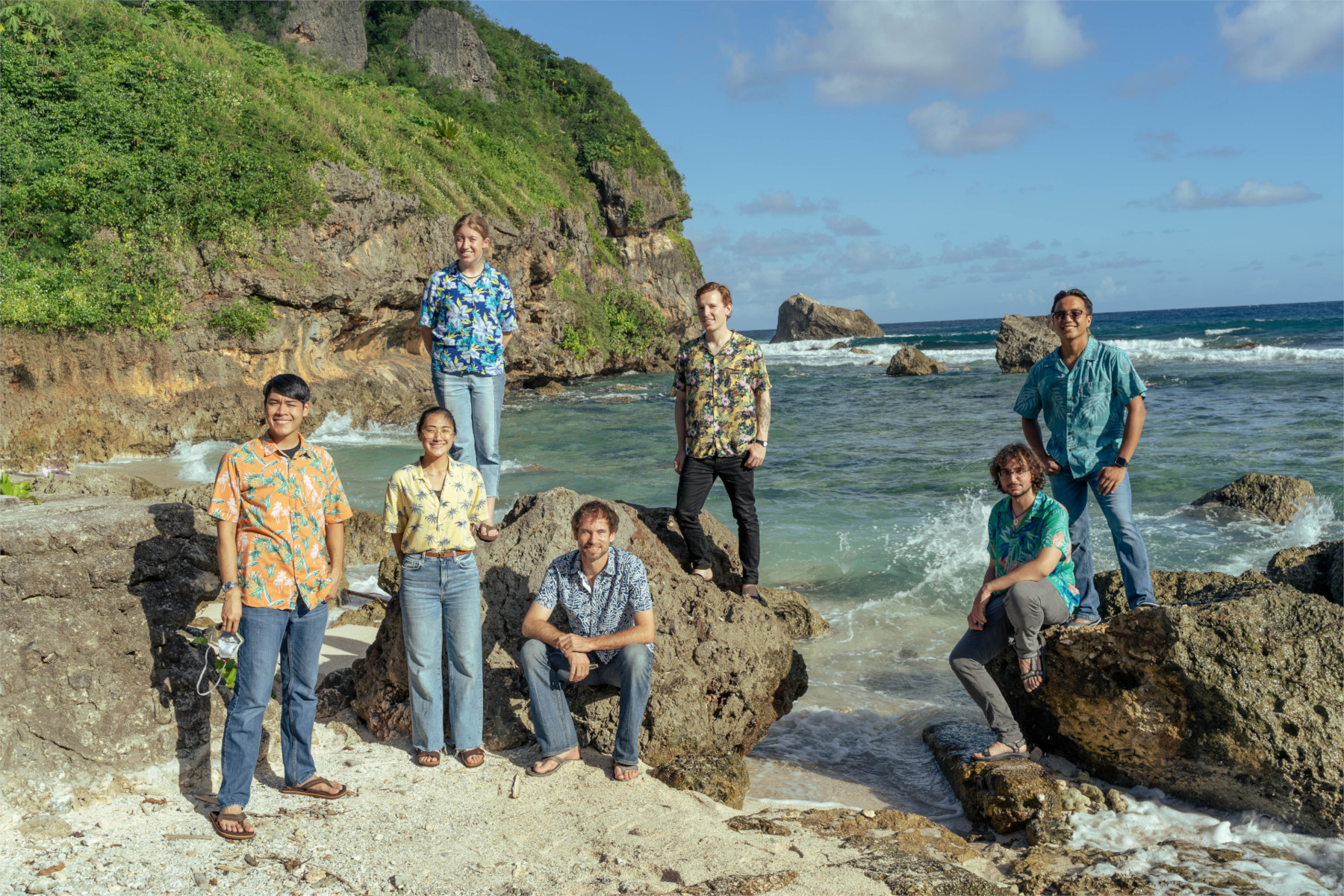

Combosch David
 Office Hours:
Office Hours:  Office Location:
Office Location: Expertise
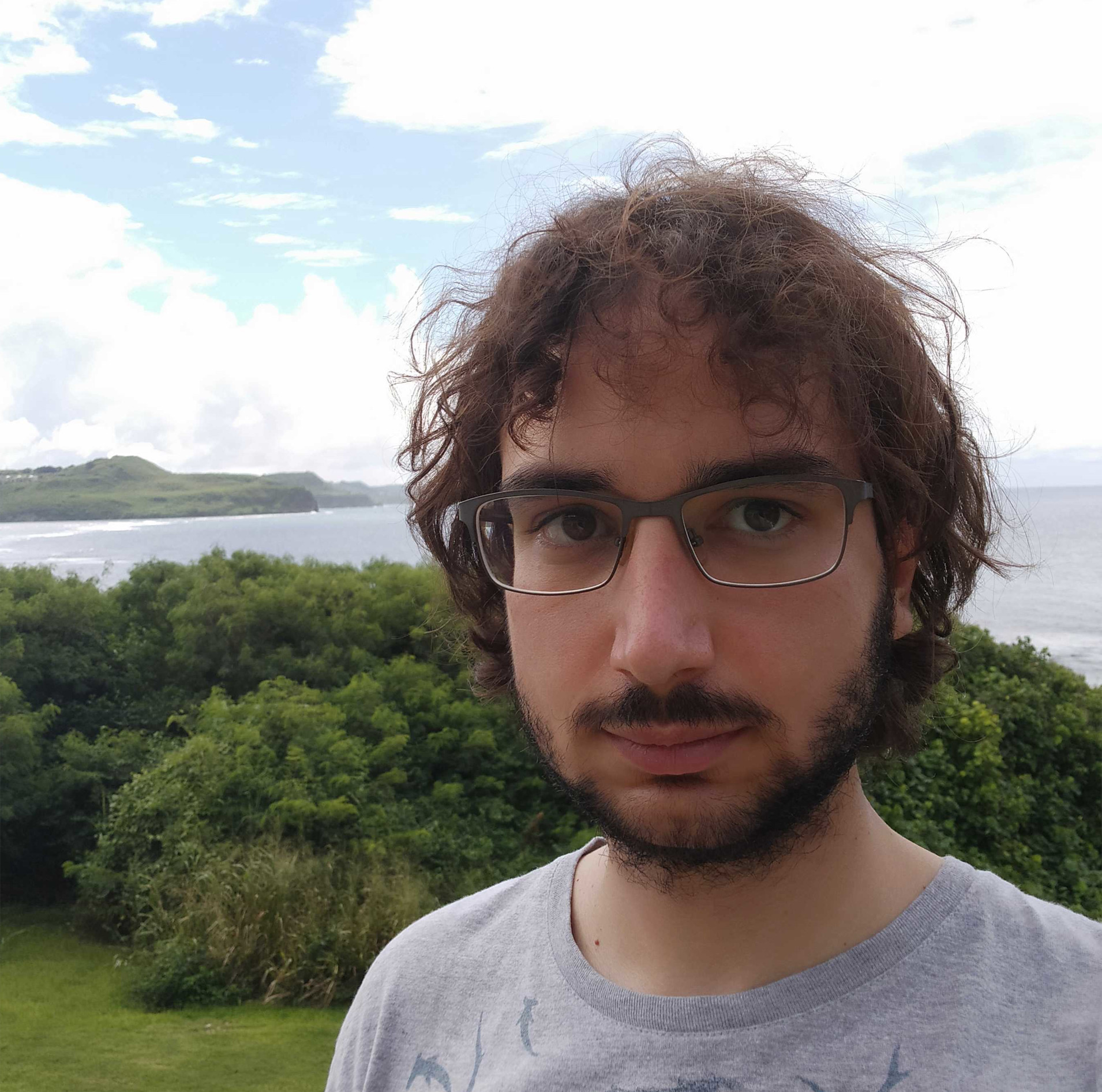

Héctor Torrado
Comunitat Valenciana, Spain
Current position: Postdoctoral Researcher
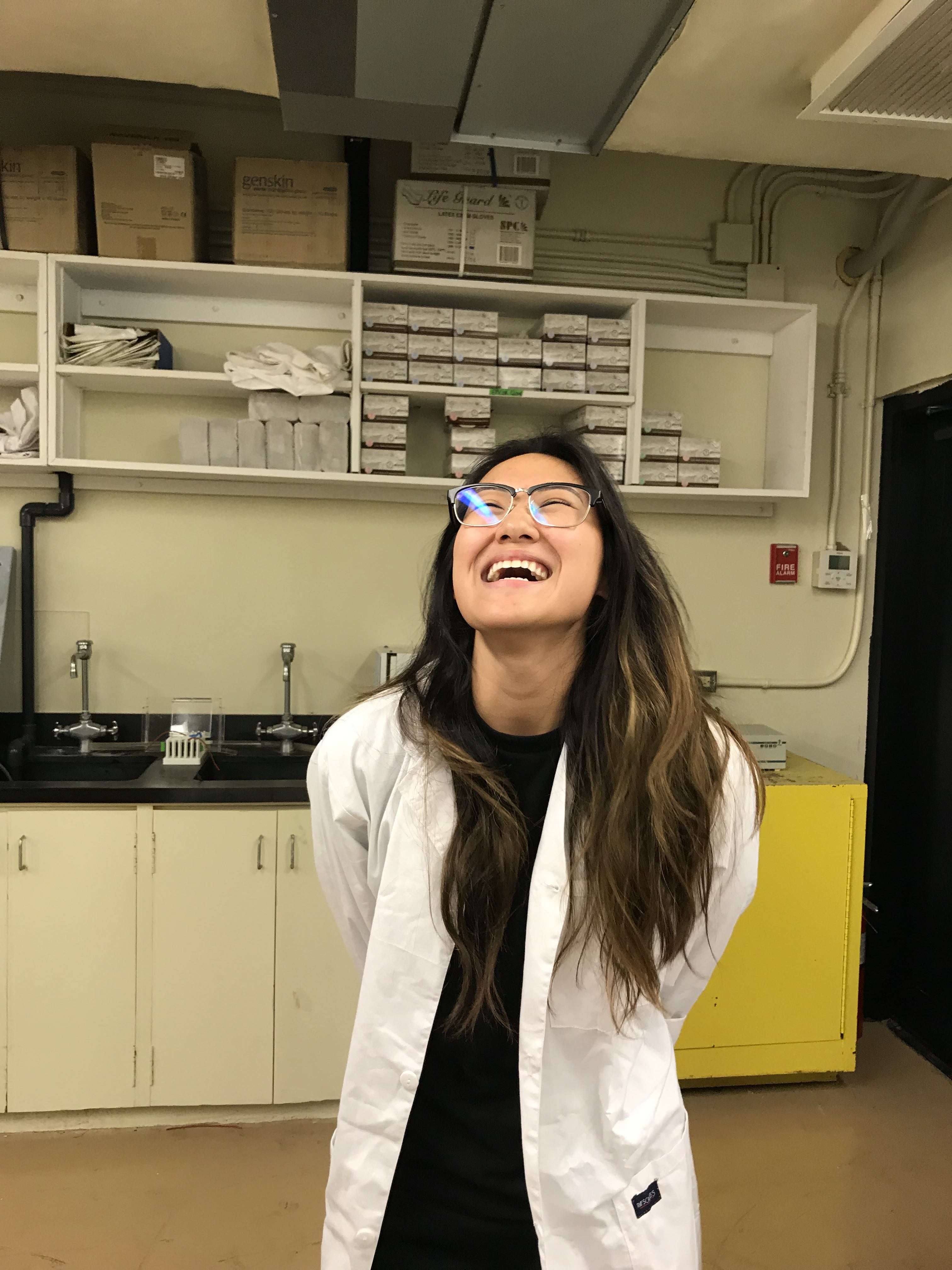

Jessica Fernandez
Yigo, Guam
Research: Coral population genomics
Current position: Graduate Student Assistant in the UOG Graduate Biology Program
About
Jess started out in the lab as a EPSCoR fellow in January 2019, working on Porites barcoding and phylogenetics. She graduated from UOG in 2020 with a major in Biology and a minor in Chemistry.Jess then re-joined the lab as grad student for the Biology Program in August 2020. She is currently working on a population genomic study of reef corals around Guam, sponsored by the NOAA Coral Reef Conservation Program.
In January 2021, Jess obtained an NSF EPSCoR fellowship for her MS thesis research.

Arthur Perez
Chalan Pago, Guam
Research: Porites phylogenomics
Current position: Graduate Student Assistant in the UOG Graduate Biology Program
Criminalist at the Guam Police Department (GPD) Crime Lab
About
Arthur received his Bachelor’s degree in Biology at the University of Guam. After graduation he worked at the Guam Museum as a data entry clerk for the Richard H. Randall Coral Collection. He also worked for the Center for Island Sustainability as a field biologist. Now he works as a criminalist at the Guam Police Department Crime Lab. He aims to gain knowledge and skills in genetics, which he can then use in human forensic DNA work.
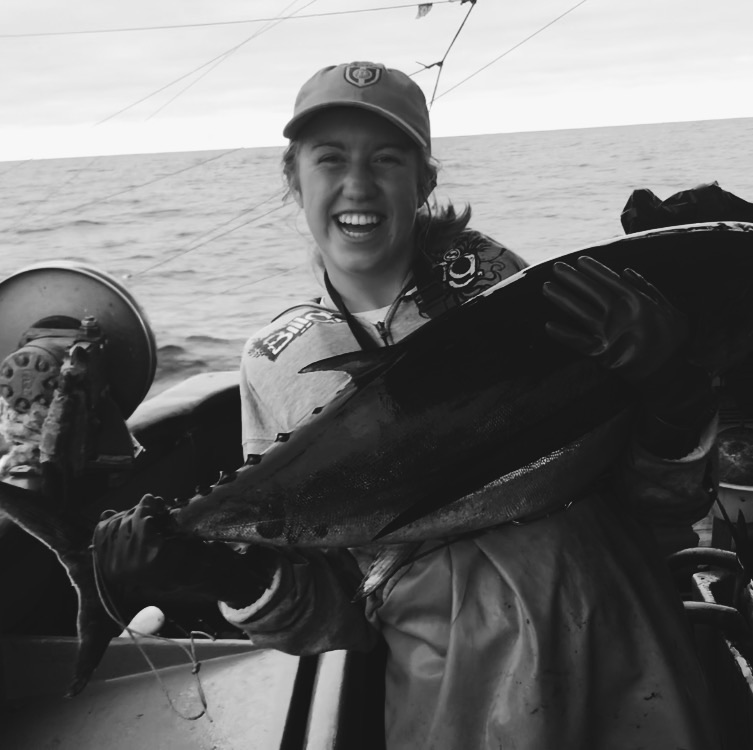

Mikay Reuter
Corvallis, Oregon
Research: Coral Restoration Genetics
Current position: Graduate Student Assistant in the UOG Graduate Biology Program
About
Mikay graduated from Oregon State University in June 2021 with a degree in Oceanography and a minor in Environmental Science. In her undergraduate degree she was able to participate in coral ecology field research in both Thailand and Indonesia. She used her coral background to work as a member of the Vega-Thurber lab helping establish a metadatabase for the Mo'orea Virus Project. In addition to 'hard science' experience, Mikay also worked as a CIMRS research fellow and 2021 Oregon Sea Grant scholar helping to improve science communication for Oregon fisheries management. She is excited to expand on these experiences and dive further into the world of genetics while researching coral restoration genetics as part of the Island Evolution Lab.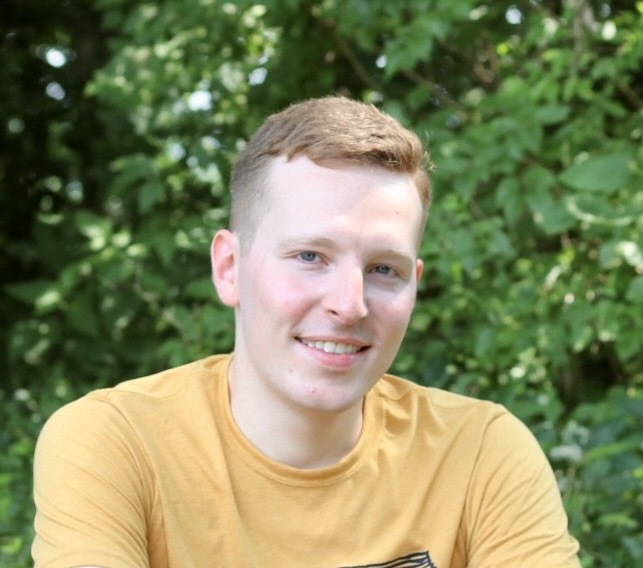
Joe Proietti
Cincinnati, Ohio
Research: Coral Population Genomics
Current position: Graduate Student Assistant in the UOG Graduate Biology Program
About
Joe graduated from Eastern Michigan University in 2018 with a BS in Environmental Biology. As an undergraduate, he worked on a prairie restoration ecology project, and also worked on a project studying the interaction between land use, nutrient pollution, and cyanobacteria blooms in a local watershed. After graduating, he worked on harmful algal bloom projects for USGS at the Great Lakes Science Center, and then worked at the NYU School of Medicine in a zebrafish developmental genetics lab. Joe joined the Island Evolution Lab in August 2021 as an NSF EPSCoR Fellow, and he is excited to work on the population genomics of corals!
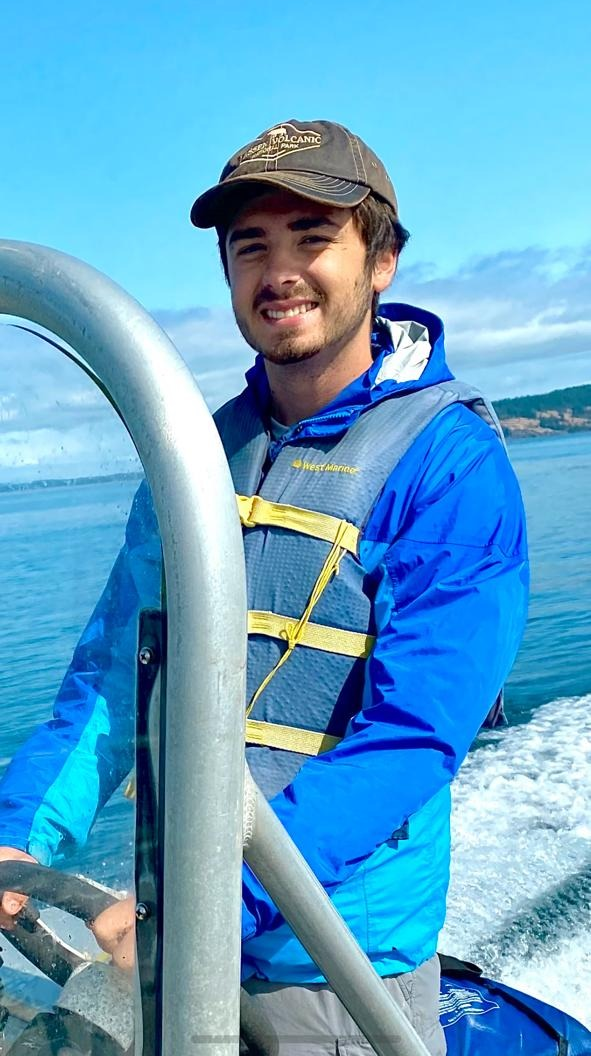

Garret O'Donnell
West Palm Beach, Florida
Research: Coral Population Genomics
Current position: Graduate Student Assistant in the UOG Graduate Biology Program
About
Garret is a Master's student studying tropical scleractinian population genetics and ecology. He studied parasitic crab diversity and curation at the Florida Museum of Natural History during his undergrad at the University of Florida. After graduating in 2021, he studied kelp forest invertebrates in Friday Harbor Washington, and soft bottom infauna communities in Fort Pierce Florida. He is interested in projects focused on stress response, diversity quantification, and specimen photography as a means to better understand the reefs of Guam and the broader pacific arena.
INTERNS:
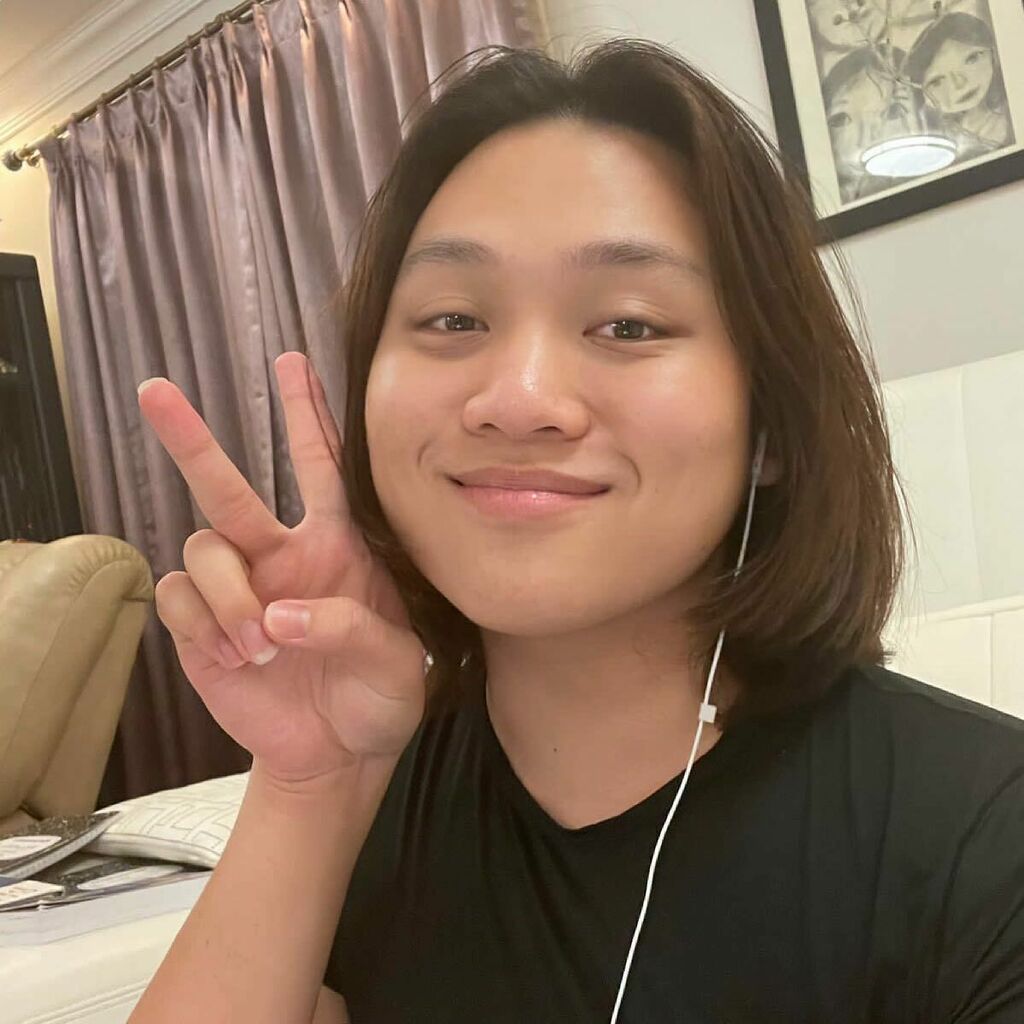

About
Felix is interested in pursuing a field in STEM and is exploring the many options in the large field. He is currently examining the morphology of several species of Staghorn corals.LAB ALUMNI:

Dareon Rios
Chalan Pago, GU
About
Dareon did her MS thesis on the population genetics of a locally dominant reef-building Acropora coral. Her research identified the population structure of this restoration target to better inform coral reef conservation and restoration management.2016 BS Human Biology, University of California San Diego, CA
2020 MS Biology, University of Guam, GU

Karim D. Primov
Miami, Florida
Research: Population Genomics of massive Porites
Current position: PhD student in the labs of Misha Matz and Mark Kirkpatrick at UT Austin
About
Karim received his Bachelor’s degree in Marine and Atmospheric Science at the University
of Miami, majoring in Marine Science and Biology and minoring in chemistry.
He studied the population genetic structure of massive Porites corals on Guam. His major research interests include population genetics and invertebrate
zoology.
Glynn, Peter W., Brian Coffman, Michael PC Fuller, Shannon G. Moorhead, Megan K. Williams, Karim D. Primov, Tayla N. Fortson, Rachel N. Barrales, and Peter J. Glynn. "Benthic ctenophores (Platyctenida: Coeloplanidae) in south Florida: environmental conditions, habitats, abundances, and behaviors." Invertebrate Biology 136, no. 4 (2017): 379-393.
Glynn, Peter W., Brian Coffman, Karim D. Primov, Shannon G. Moorhead, Jeongran Vanderwoude, Rachel N. Barrales, Megan K. Williams, and Robert P. Roemer. "Benthic ctenophores (Platyctenida: Coeloplanidae) in South Florida: predator–prey interactions." Invertebrate Biology (2018).

Leya Wang
Mangilao, Guam
Research: Pocillopora reproduction
Bio: High School Intern (2018-2019); Senior at St. Johns High School, Tumon, Guam
About
Leya graduated from St. Johns High School in June 2019 and started college at John Hopkins University in the Fall semester 2019.
Kireon Rios
Chalan Pago, Guam
Research: Porites barcoding and phylogenetics
Current position: Undergraduate Intern;
Senior at University of Guam, Biology Program
CONGRATULATIONS Kireon for receiving the Louis Stokes Alliances for Minority Participation (LSAMP) stipend for his research on Porites barcoding and phylogenetics.
About
Kireon joined the lab in January 2019 as an undergraduate intern working on Porites barcoding and phylogenetics.10/2019: Kire just received the Louis Stokes Alliances for Minority Participation (LSAMP) stipend for his research on Porites barcoding and phylogenetics.

Christian Binuya
Yigo, GU
Research: Pocillopora barcoding and phylogenetics
Current position: Senior at University of Guam, double majoring in chemistry and biology under the Chemistry Program
About
01/2020: Christian received the EPSCoR (NSF) undergraduate research stipend for his research on Pocillopora barcoding and phylogenetics.Why I chose to be part of the team: I want to gain better understanding of how genetics research is used not only in the medical field, but also in studying evolution and ecology.
Research of interests: ultimately, I am interested in two completely different fields; the field of applied genetics research, and conservation research.
Career goal: become a physician and continue a career in research (MD program, MD/PhD program) in hoping to give back to the community in Guam.

Andrea Fuentes
University Park, IL
Advisor: Dr. David Combosch
Research: Porites Coral Phylogenetics
Bio: Visiting Undergraduate Student (2018) and back then Senior at Governors State University, IL
About
Andrea graduated from Governors State University, IL, in June 2019Main Team Collaborations:
- Gonzalo Giribet (Harvard University, Cambridge)
- Ana Riesgo (Museum of Natural History, London)
- Neil Landman (American Museum of Natural History, NY)
- Martin Schwentner (Natural History Museum, Vienna)
- Valerie Paul (Smithsonian National Museum of Natural History)
- Rüdiger Bieler (Field Museum, Chicago)
- Sarah Davies (Boston University)
- Sarah Lemer (University of Guam)
- Laurie Raymundo (University of Guam)
- Mike Sweet (University of Derby)
- Dave Baker (University of Hong Kong)
- Crawford Drury (University of Hawaii)
- Bernardo Lemos (Harvard University, Boston)
- Miquel Arnedo (University of Barcelona)
- Rich Aronson (Florida Institute of Technology)
- Lauren Toth (US Geological Survey)
- Zac Forsman (University of Hawaii)
- Dan Barshis (Old Dominion University, Norfolk)
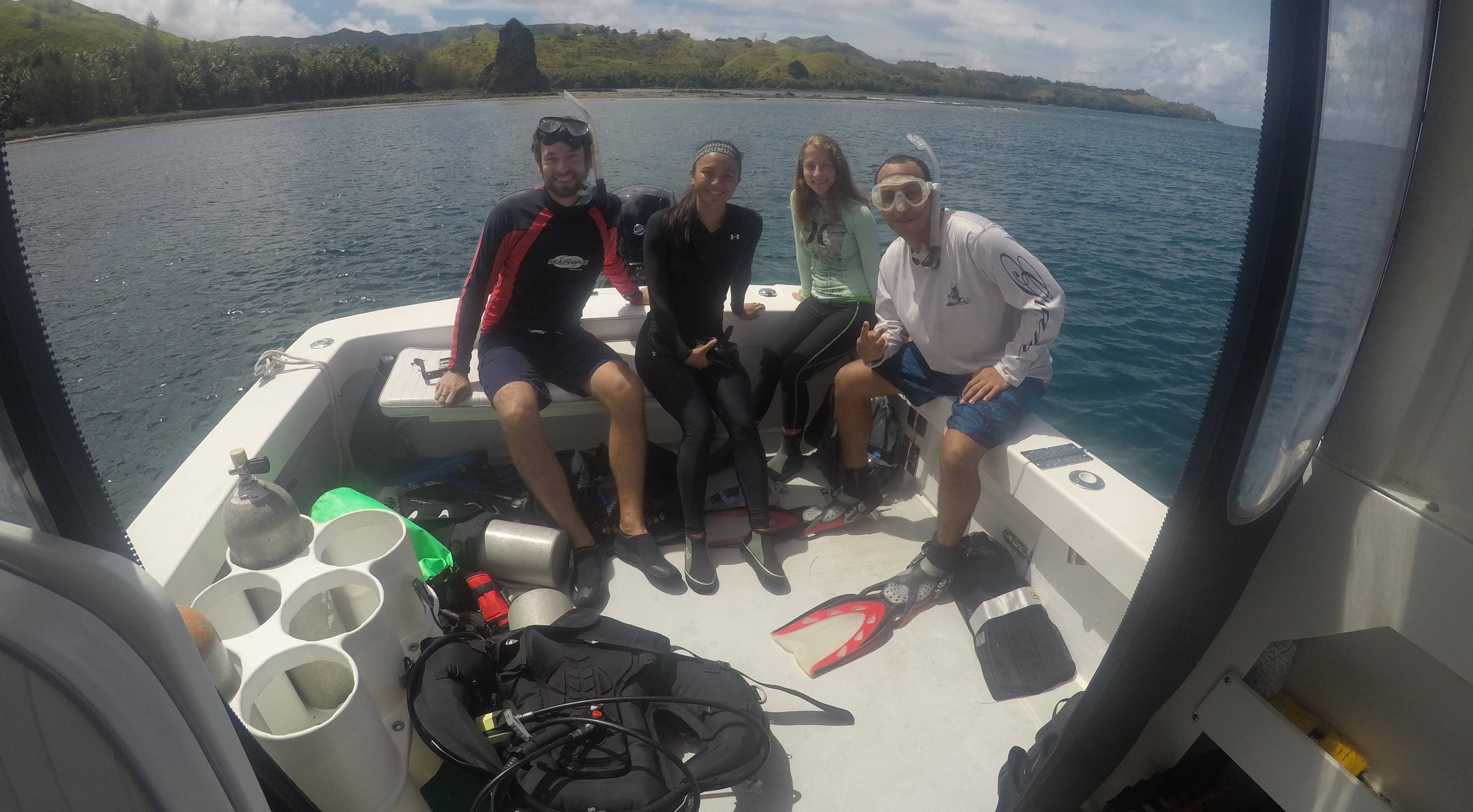
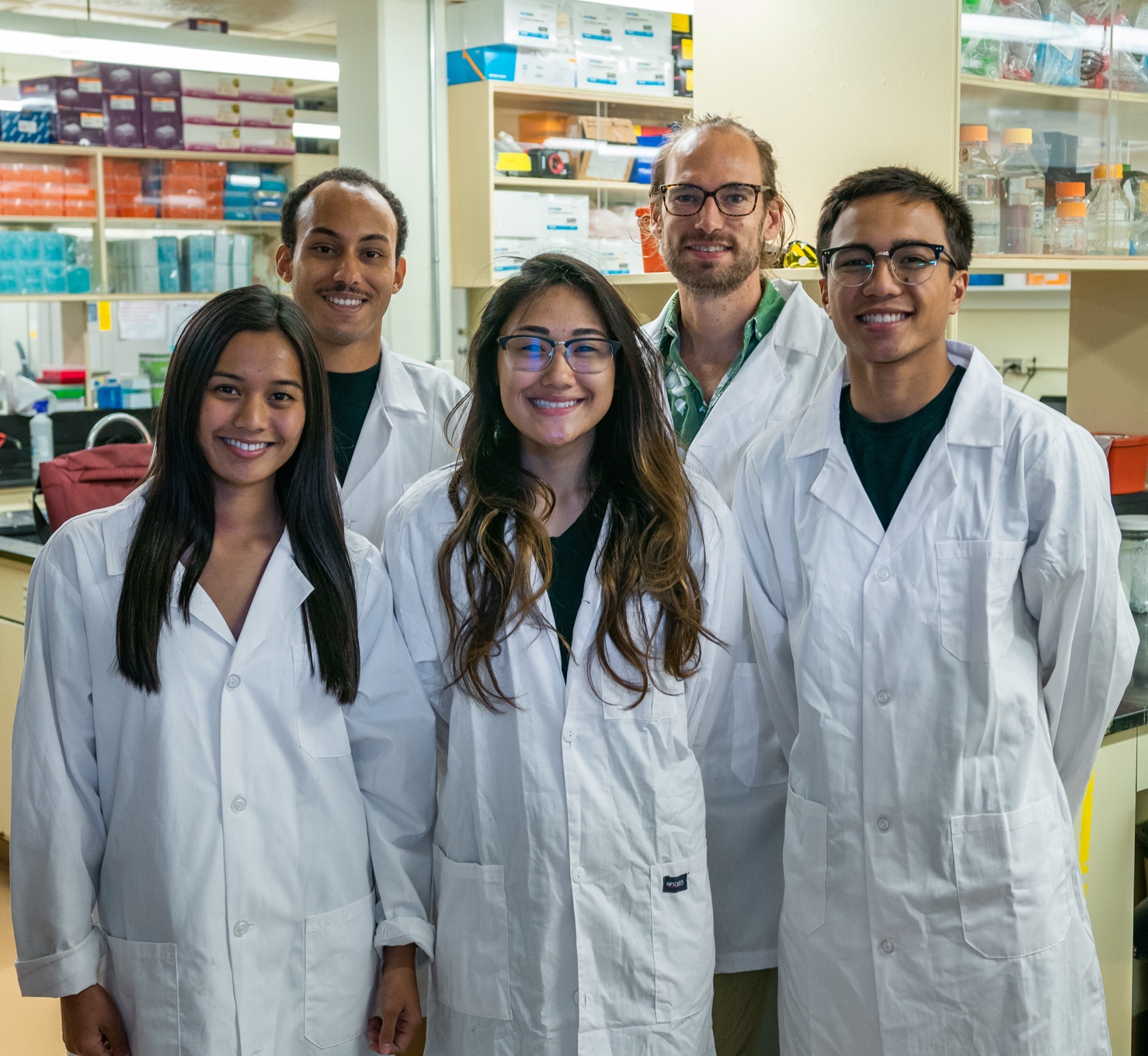
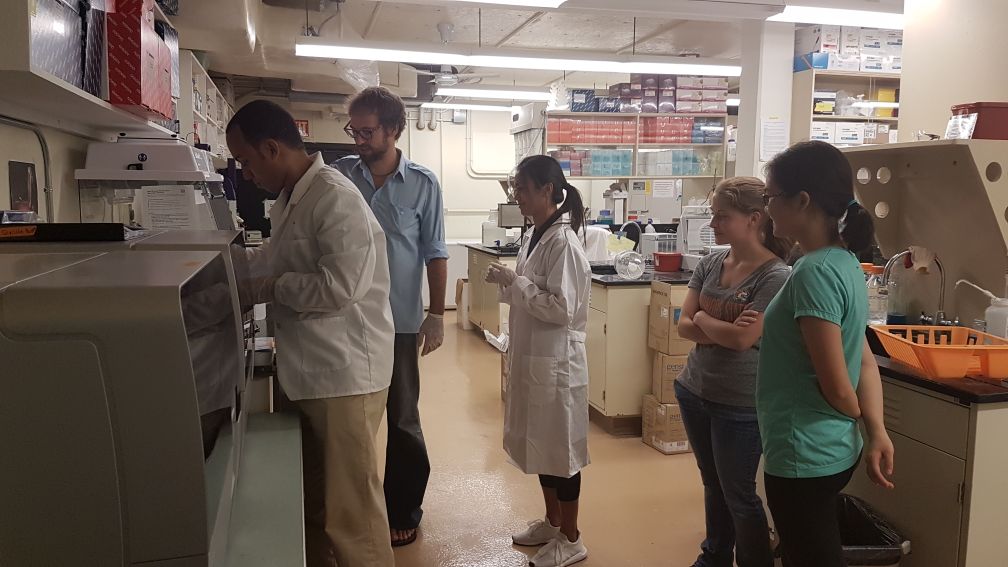
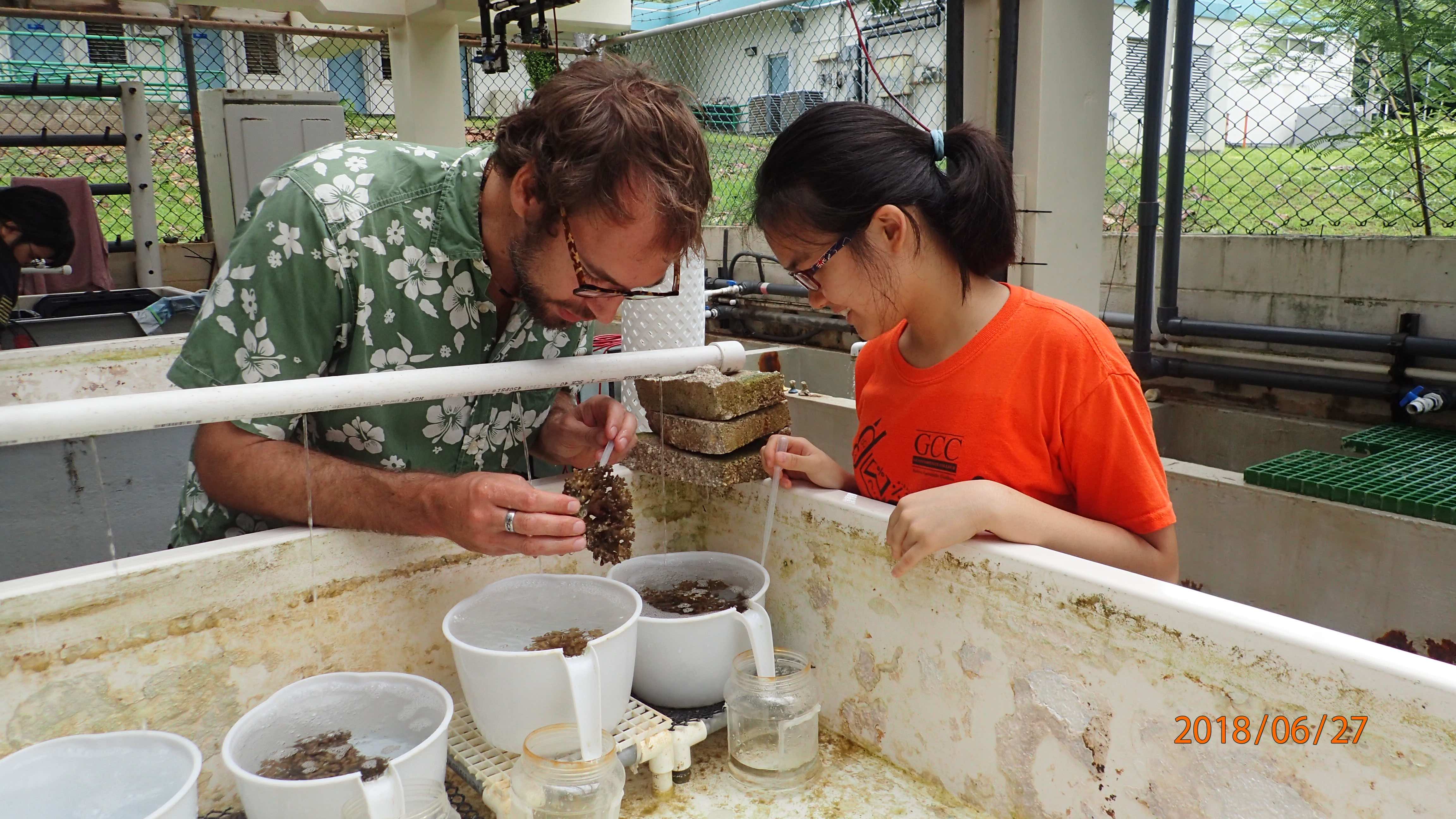




Coral Population Genomics
We are exploring the genetic connectivity, diversity and structure of multiple coral species around Guam and throughout Micronesia using an in-house RAD-Seq approach.
This is a major focus of our lab and most graduate sutdent thesis (e.g. Dareon, Karim and Joe) focus on coral population genomics (see team).
Coral Phylogenomics & Phylogenetics
The main phylogenetic relationships among corals have been highly controversial for years. Recently genome-sclae datasets have started to clarify many of these important relationships. We are currently assembling on one of the largest genomic dataset to study the deep relationships in the coral tree of life. In addition, we are working on a phylogenetic barcoding study of Porites corals.
Coral Bleaching Resistance
We are using experiments to induce coral bleaching in aquaria experiment to characterize the molecular processes that allow corals to survive and adapt to stressful environmental conditions. Most of these projects are conducted in collaboration with the Lemer Lab.
Nautilus Phylogeography
The Nautilus is one of the most iconic marine invertebrates and an important model system for evolutionary biologist and paleontologists, among others. We are using genomic tools to analyze the evolution and phylogeography of Nautilus across the species distribution range. We currently work on the discriptions of several new Nautilus species based on Combosch et al 2017.
Coral Reproduction
Coral reproduction is of major interest, not only to improve our understanding of the basic biology of reef corals but also for their management and restoration. We are studying the reproductive characteristics of several coral species, using aquarium experiments and genetic tools.
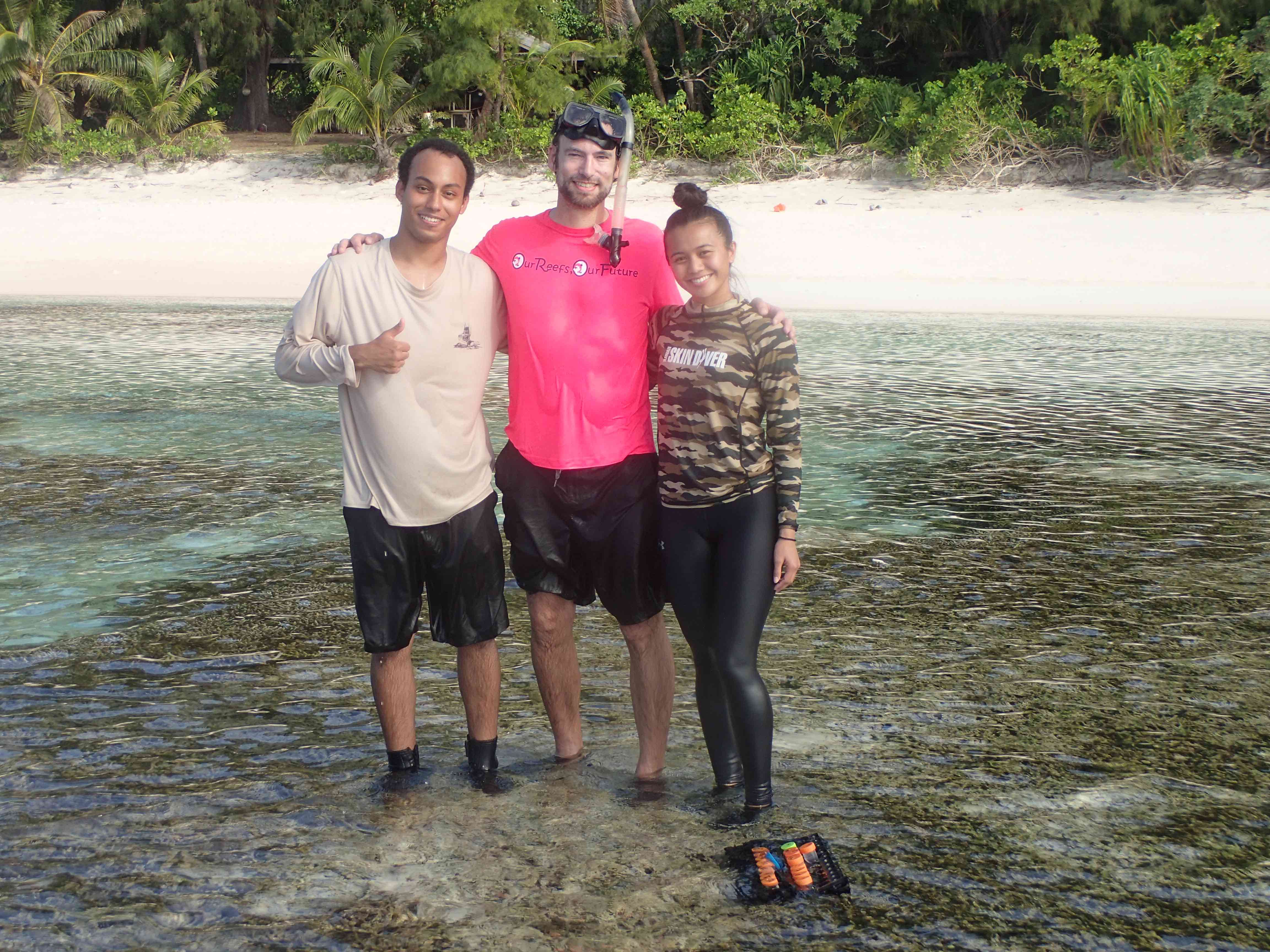
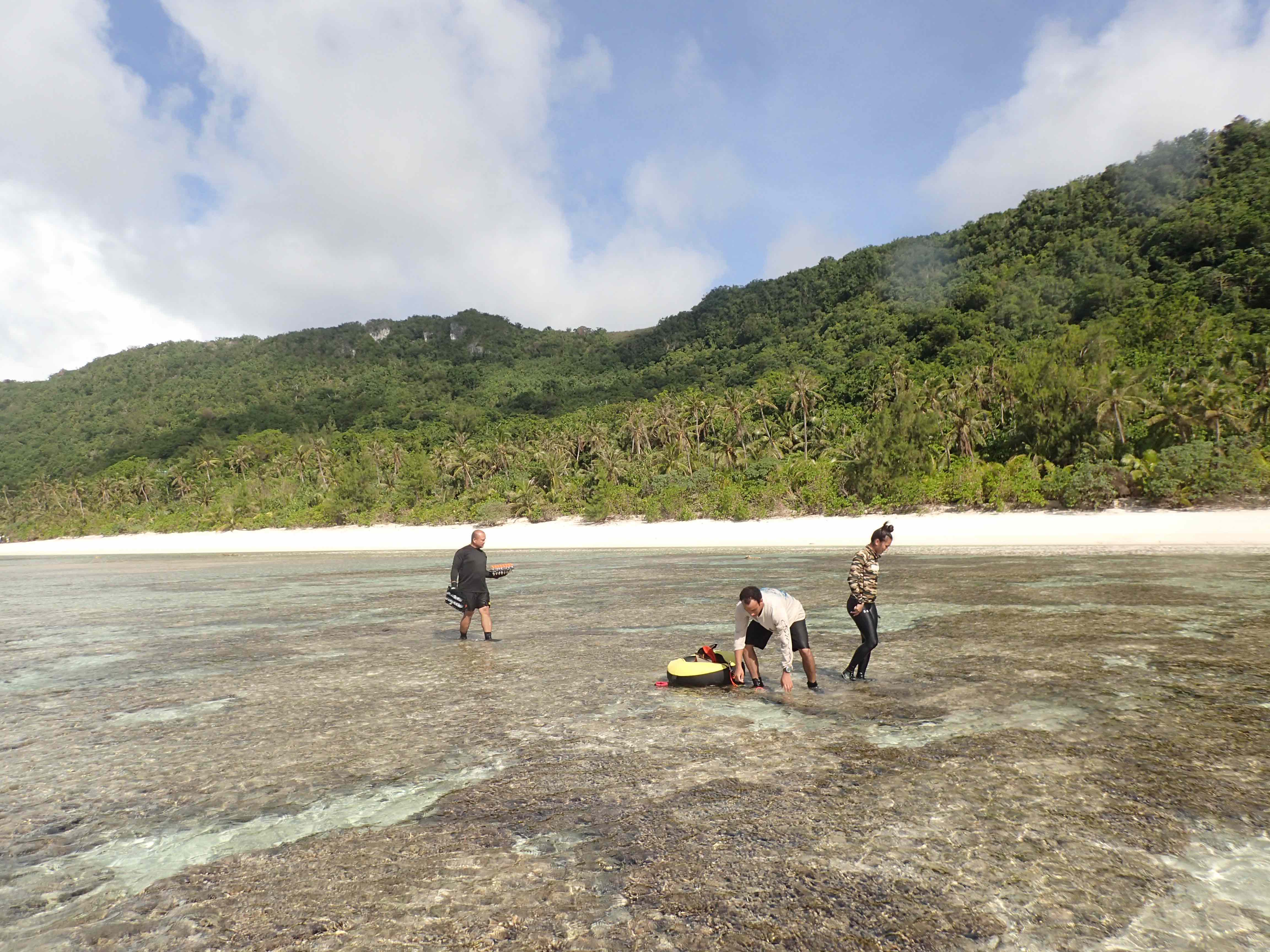
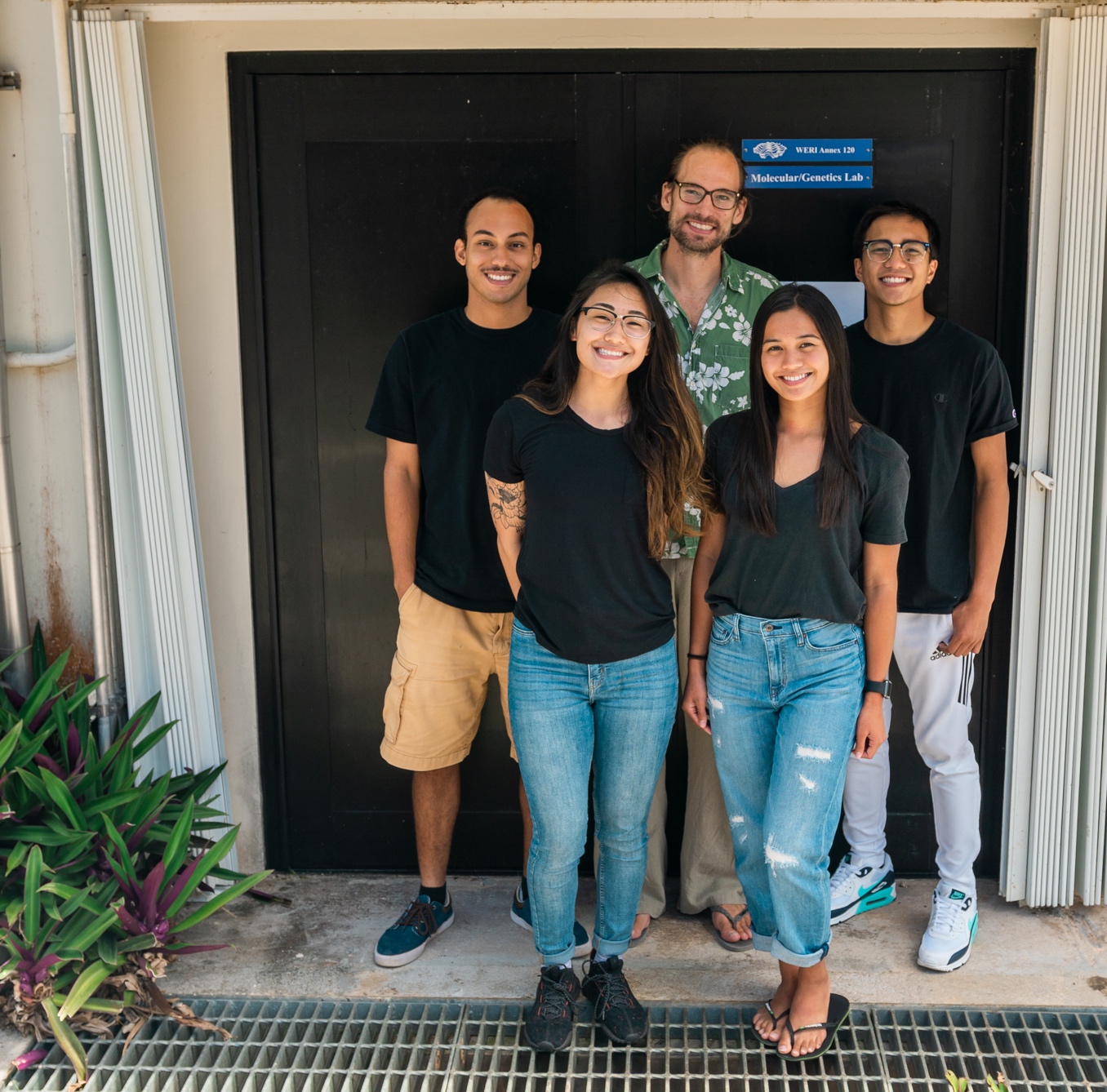






David's Google Scholar Profile
Please contact us for pdf copies or download from researchgate or sci-hub.
Select Publications:
2021
- Voolstra C. R., K.M. Quigley, S.W. Davies, J.E. Parkinson, R.S. Peixoto, M. Aranda, A.C. Baker, A.R. Barno, D.J. Barshis, F. Benzoni, V. Bonito, D.G. Bourne, C. Buitrago-López, T.C.L. Bridge, C.X. Chan, D.J. Combosch, J. Craggs, J.C. Frommlet, S. Herrera, A.M. Quattrini, T. Röthig, J.D. Reimer, E. Rubio-Portillo, D.J. Suggett, H. Villela, M. Ziegler & M. Sweet (2021) Consensus guidelines for advancing coral holobiont genome and specimen voucher deposition. Frontiers in Marine Science 8:1029.
- Moles J., T.J. Cunha, S. Lemer, D.J. Combosch & G. Giribet 2021 Tightening the girdle: Phylotranscriptomics of Polyplacophora. Journal of Molluscan Studies 87(2): eyab019
2020
- Schwentner M., G. Giribet, D.J. Combosch, & B.V. Timms 2020 Genetic Differentiation in Mountain-Dwelling Clam Shrimp, Paralimnadia (Crustacea: Branchiopoda: Spinicaudata), in Eastern Australia. Invertebrate Systematics 34 (1): 88–100. doi:10.1071/IS19027.
2019
- Walker N.S., R. Fernández, J.M. Sneed, V.J. Paul, G. Giribet, & D.J. Combosch 2019 Differential Gene Expression During Substrate Probing in Larvae of the Caribbean Coral Porites astreoides. Molecular Ecology 28(22): 4899-4913, DOI:10.1111/mec.15265.
- Leiva C., S. Taboada, N.J. Kenny, D. Combosch, G. Giribet, T. Jombart, A. Riesgo (2019) Population substructure and signals of divergent adaptive selection despite admixture in the sponge Dendrilla antarctica from shallow waters surrounding the Antarctic Peninsula. Molecular Ecology 28(13): 3151-3170, DOI: 10.1111/mec.15135
- Laumer C., R. Fernández, S. Lemer, D. Combosch, K. Kocot, A. Riesgo, S. Andrade, W. Sterrer, M. Sørensen, G. Giribet (2019) Revisiting metazoan phylogeny with genomic sampling of all phyla. Proceedings of the Royal Society B. 286(1906): 20190831, DOI: 10.1098/rspb.2019.0831
2017
- Combosch* D.J., S. Lemer*, P.D. Ward, N.H. Landman & G. Giribet 2017 Genomic signatures of evolution in Nautilus–an endangered living fossil. Molecular Ecology 26(21): 5923–5938 *Shared first author
- Schwentner M., D.J. Combosch, J.P. Nelson & G. Giribet 2017 A Phylogenomic Solution to the Origin of Insects by Resolving Crustacean-Hexapod Relationships. Current Biology 27(12): 1818-1824
- Combosch D.J., T.M. Collins, E.A. Glover, D.L. Graf, E.M. Harper, J.M. Healy, G.Y. Kawauchi, S. Lemer, E. McIntyre, E.E. Strong, J.D. Taylor, J.D. Zardus, P.M. Mikkelsen, G. Giribet & R. Bieler 2017 A family-level Tree of Life for bivalves based on a Sanger-sequencing approach. Molecular Phylogenetics & Evolution 107: 191–208
2016
- Combosch D.J. & G. Giribet 2016 Clarifying phylogenetic relationships and the evolutionary history of the bivalve order Arcida (Mollusca: Bivalvia: Pteriomorphia). Molecular Phylogenetics & Evolution 94: 298-312
2015
- Combosch D.J. & S.V. Vollmer 2015 Trans-Pacific RAD-Seq population genomics confirms introgressive hybridization in Eastern Pacific Pocillopora corals. Molecular Phylogenetics & Evolution 88: 154-162
2013
- Combosch D.J. & S.V. Vollmer 2013 Mixed asexual and sexual reproduction in the Indo-Pacific reef coral Pocillopora damicornis. Ecology & Evolution 3(10): 3379-3387
2012
- Toth L.T., R.B. Aronson, S.V. Vollmer, J.W. Hobbs, D. Urrego, H. Cheng, I. Enochs, D.J. Combosch, R. van Woesik & I. Macintyre 2012 ENSO drove 2500-Year Collapse of Eastern Pacific Coral Reefs. Science 337(6090): 81-84
2011
- Combosch D.J. & S. Vollmer 2011 Population Genetics of an Ecosystem-Defining Reef Coral Pocillopora damicornis in the Tropical Eastern Pacific. PLoS ONE 6(8): e21200
2008
- Combosch D.J., H. Guzmán, H. Schuhmacher & S. Vollmer 2008 Interspecific Hybridization and Restricted Gene Flow in Tropical Eastern Pacific Pocillopora. Molecular Ecology 17: 1304-1312
Presentations
2019
- Asia Pacific Academy of Science, Education and Environmental Management (APASEEM),
Garapan, Saipan (CNMI)
- Combosch D.J.: Population genomics of reef-building corals in the CNMI and Guam. -
Gordon Research Conference: Marine Molecular Ecology, Hong Kong
- Combosch D.J.: Population genomics of reef corals in Guam. -
5th Guam Coral Reef Symposium, Tumon (Guam)
- Rios D.* & D.J. Combosch: Population Genomics of Acropora pulchra in Guam.
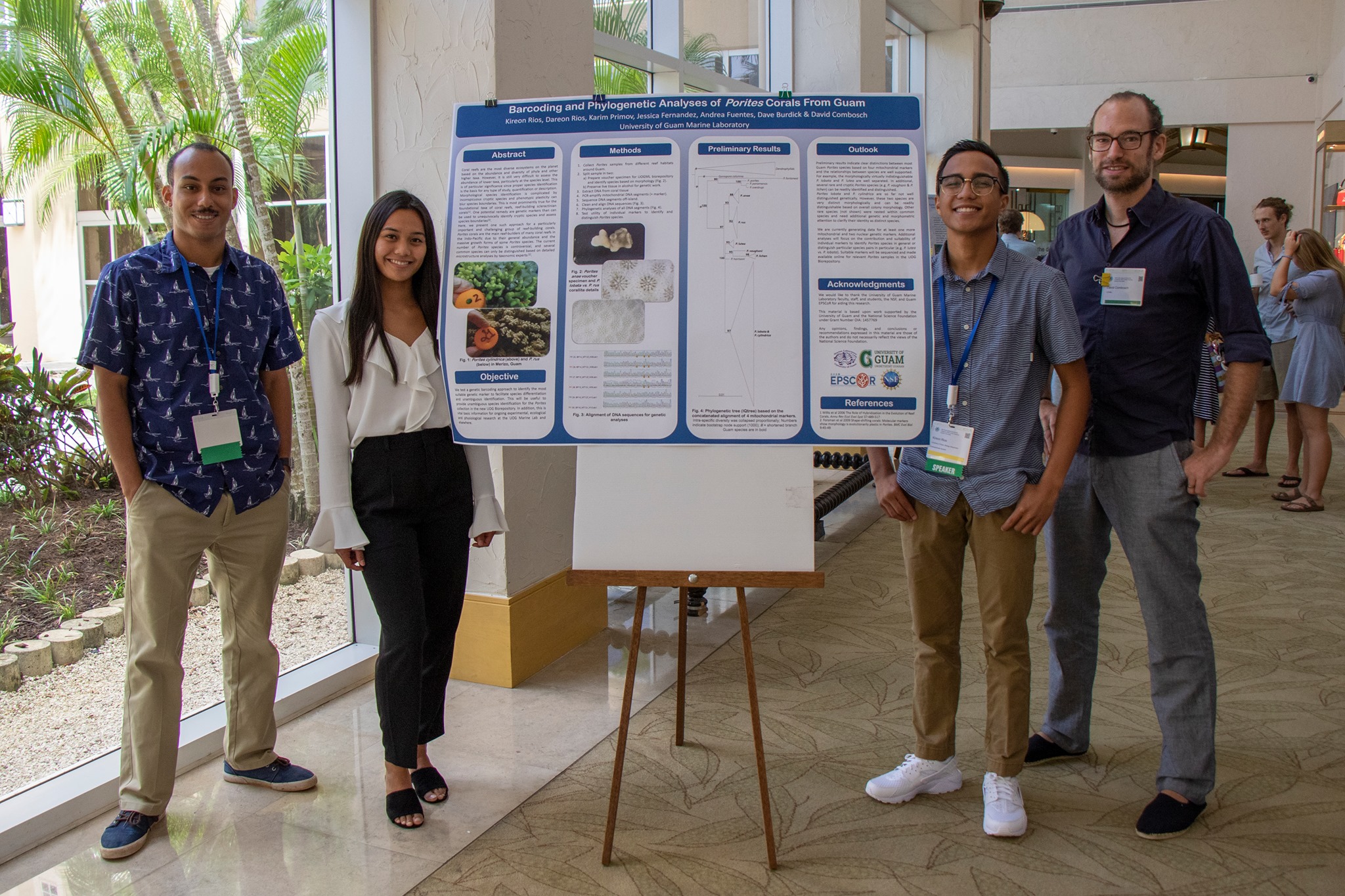
- Rios K.**, D. Rios, K. Primov, J. Fernandez, A. Fuentes, D. Burdick & D.J. Combosch: Barcoding and phylogenetic analyses of reef-building Porites corals.
- Primov K.* & D.J. Combosch: The Population Genetic Structure of the reef-building coral Porites lobata on Guam. -
CISX – 10th Conference for Island Sustainability, Tumon (Guam)
- Rios D.* & D.J. Combosch: The Population Genetic Structure of Acropora pulchra in Guam.
- Primov K.* & D.J. Combosch: Population Genetic Structure of the reef-building coral Porites lobata on Guam.
2018
- University of Guam (Guam), POETS Seminar Series
- Combosch D.J.: Genomic signatures of evolution in Nautilus – an endangered living fossil. -
SACNAS - The National Diversity in STEM Conference, San Antonio, TX (USA)
- Fuentes A.** & D.J. Combosch: Resolving the Species Boundaries in the Porites Corals of Guam. -
Willi Henning Congress, Barcelona (Spain)
- Enguidanos A.*, D.J. Combosch, V. Tonzo, G. Giribet & M. Arnedo: Knocking on the trap-door: Unraveling the species boundaries and evolutionary history of western Mediterranean ctenizid trap-door spiders. -
Asia-Pacific Coral Reef Symposium, Cebu (Philippines)
- Combosch D.J., S. Lemer, R. Bieler & G. Giribet: A genome and transcriptome-based phylogeny of reef corals. -
4th Guam Coral Reef Symposium, Tumon (Guam)
- Combosch D.J., S. Lemer, R. Bieler & G. Giribet: A phylogenomic study of Scleractinian corals based on genome and transcriptome data. -
CISX – 9th Conference for Island Sustainability, Tumon (Guam)
- Primov K.* & D.J. Combosch: The Population Genetic Structure of Porites lobata on Guam. -
Connections, Collaborations and Innovations in the Pacific, UOG Annual Research Conference, Mangilao (Guam)
- Combosch D.J.: Connections, Collaborations and Innovations and the EPSCoR Guam Ecosystem Collaboratorium.
2017
- Hong Kong University (Hong Kong), Ecology and Biodiversity Seminar
- Combosch D.J. & S. Lemer: Evolutionary Genomics and the EPSCoR Guam Ecosystem Collaborative. - University of Guam (Guam), SECORE Coral Reproduction workshop
- Combosch D.J.: Population Genetics and Restoration. - International Society for Reef Studies European Meeting, Oxford (UK)
- Combosch D.J., S. Lemer, R. Bieler & G. Giribet: A genome and transcriptome based phylogenomic study of Scleractinian corals.
- Lemer S., D.J. Combosch & B. Lemos: Epigenetic modification and differential gene expression in Acropora digitifera during heat acclimation. - 10th World Sponge Conference, Galway (Ireland)
- Leiva C.*, S. Taboada, D.J. Combosch, G. Giribet & A. Riesgo: Genomic approach to the evolutionary history and population structure of Dendrilla antarctica Topsent, 1905 (Porifera, Demospongiae) from the shallow-water Southern Ocean. - XIIth Scientific Committee on Antarctic Research, Leuven (Belgium)
- Taboada S., C. Leiva, G. Giribet, D.J. Combosch, J. Junoy, J. Avila & A. Riesgo: Molecular connectivity in two congeneric Antarctic shallow-water hoplonemertean species with direct development using a traditional marker and SNPs
- Leiva C.*, S. Taboada, D.J. Combosch, G. Giribet & A. Riesgo: Population connectivity of Dendrilla antarctica Topsent, 1905 (Porifera, Demospongiae) from West Antarctica shallow bottoms using genome-wide SNPS obtained from RADSeq techniques.
2016
- 12th International Coral Reef Symposium, Honolulu, HI (USA)
- Combosch D.J., S. Lemer & G. Giribet: A phylogenomic backbone for Scleractinia. - Harvard University OEB Thesis Symposium, Cambridge, MA (USA)
- Walker N.*, G. Giribet & D.J. Combosch: Characterizing Differential Gene Expression in Probing Larvae of the Caribbean Coral Species Porites astreoides Lamarck, 1816.
2015
- Harvard Museum of Comparative Zoology (USA), MCZ Seminar Series
- Combosch D.J.: Population genomics of nautilids. - Gordon Research Conference: Ecological and Evolutionary Genomics, Biddeford, ME (USA);
- Combosch D.J., S. Lemer & G. Giribet: Population genomics of nautilids.
2012
- 12th International Coral Reef Symposium, Cairns (Australia)
- Combosch D.J. & S.V. Vollmer: Transpacific phylogenomic analysis of Pocillopora damicornis populations.
2010
- International Society for Reef Studies European Meeting, Wageningen (Netherlands)
- Combosch D.J. & S.V. Vollmer: Population genetics of an ecosystem-engineer - Pocillopora damicornis (Cnidaria, Scleractinia) in the Eastern Pacific.
2009
- Ruhr-University Bochum (Germany), Biodiversity Colloquium
- Combosch D.J.: Molecular Ecology of Tropical Eastern Pacific Corals. - Western Society of Naturalists, Monterey, CA (USA)
- Combosch D.J. & S.V. Vollmer: Population genetics of an ecosystem-defining reef builder - Pocillopora damicornis in the Tropical Eastern Pacific.
2008
- 11th International Coral Reef Symposium, Fort Lauderdale, FL (USA)
- Combosch D.J., H. Schuhmacher, H. Guzman & S.V. Vollmer: Local Interspecific Hybridization and Restricted Transpacific Gene Flow in Tropical Eastern Pacific Pocillopora. - 37th Benthic Ecology Meeting, Providence, RI (USA)
- Combosch D.J., H. Schuhmacher, H. Guzman & S.V. Vollmer: Interspecific Hybridization in Eastern Pacific Pocillopora (Scleractinia).
2006
- International Society for Reef Studies European Meeting, Bremen (Germany)
- Combosch D.J., H. Schuhmacher & S.V. Vollmer: Unexpected variation in the Calmodulin gene of corals.
*Student presenter; **Undergraduate presenter



Teaching:
- Evolutionary Biology (BI/EV525)
- Population Genetics Seminar (BI/EV691)
- Evolution in Changing Seas Guam
- Phenotypic Plasticity Seminar (BI/EV691)

Outreach examples: